
Diana Arseni from the MRC Laboratory of Molecular Biology, UK
Diana graduated with an MSci in Biomedicine from the University of Lancaster in 2016. During her PhD at the University of Glasgow and Astrazeneca, she investigated mechanisms through which microglia mediate tissue damage in the context of multiple sclerosis. In 2020, she started her postdoc at the MRC LMB, in the lab of Ben Ryskeldi-Falcon, where her work focused on elucidating the structure of TDP-43 filaments from neurodegenerative diseases using cryo-electron microscopy. In November 2024, she started her own research group in the Neurobiology Division at the MRC LMB. Her group studies mechanisms of brain health, ageing and disease focusing on the lysosomal protein transmembrane protein 106B (TMEM106B) which forms amyloids in the brain in an age-dependent manner.
Talk Abstract
The ABC of TDP-43
Abnormal assemblies of TDP-43 in neurons and glia are the pathological hallmark of amyotrophic lateral sclerosis (ALS) and multiple types of frontotemporal lobar degeneration (FTLD). Mutations in the TDP-43 gene, TARDBP, can cause ALS and FTLD,and the temporospatial accumulation of TDP-43 assemblies correlates with neurodegeneration, indicating a causative role for TDP-43 assembly in disease. TDP-43 assemblies are also common co-pathologies in other diseases, including Alzheimer’s, Parkinson’s and Huntington’s. The structural and molecular mechanisms of TDP-43 assembly in disease are poorly understood. We developed a protocol to isolate assembled TDP-43 from the brains of patients with ALS and FTLD and determined their structures using cryo-electron microscopy (cryo-EM). We found that TDP-43 assembles into amyloid filaments in these diseases. The ordered filament cores are comprised of the first half of the TDP-43 low-complexity domain and adopt distinct filament folds in different neurodegenerative conditions. These brain-derived filament folds show no similarity to TDP-43 filament folds formed in vitro. The structures, in combination with mass spectrometry, led to the identification of two new post-translational modifications of assembled TDP-43, citrullination and mono-methylation of R293, and suggest that they may facilitate filament formation and observed structural variation within individual filaments. Unexpectedly, the structures also revealed that in specific cases TDP-43 can also co-assemble with Annexin A11 in heteromeric amyloid filaments. The structures of TDP-43 amyloid filaments from ALS and FTLD guide mechanistic studies of TDP-43 assembly, as well as the development of diagnostic and therapeutic compounds for TDP-43 proteinopathies.

Morgan Beeby from the Imperial College London, UK
Morgan studied for a BSc at the University of Birmingham. A summer project refining a crystal structure inspired a move to a PhD under Todd O. Yeates at UCLA where he trained in crystallography and bioinformatics to understand how hyperthermophiles have adapted to survive at high temperatures, and the genomic basis for self-assembly of bacterial microcompartments and analogous structures. To better understand principles of biological self-assembly and evolution, he next trained in electron cryo-tomography with Grant J. Jensen at Caltech, working on projects including cataloging the diversity of bacterial flagellar motors, nanoscale motors that spin helical filaments to act as propellors. He started his own lab in 2013 using electron cryo-tomography as the core technique to continue work that uses the structural diversity of flagellar motors to shed light upon basic principles of molecular assembly, function, and evolution.
Talk Abstract
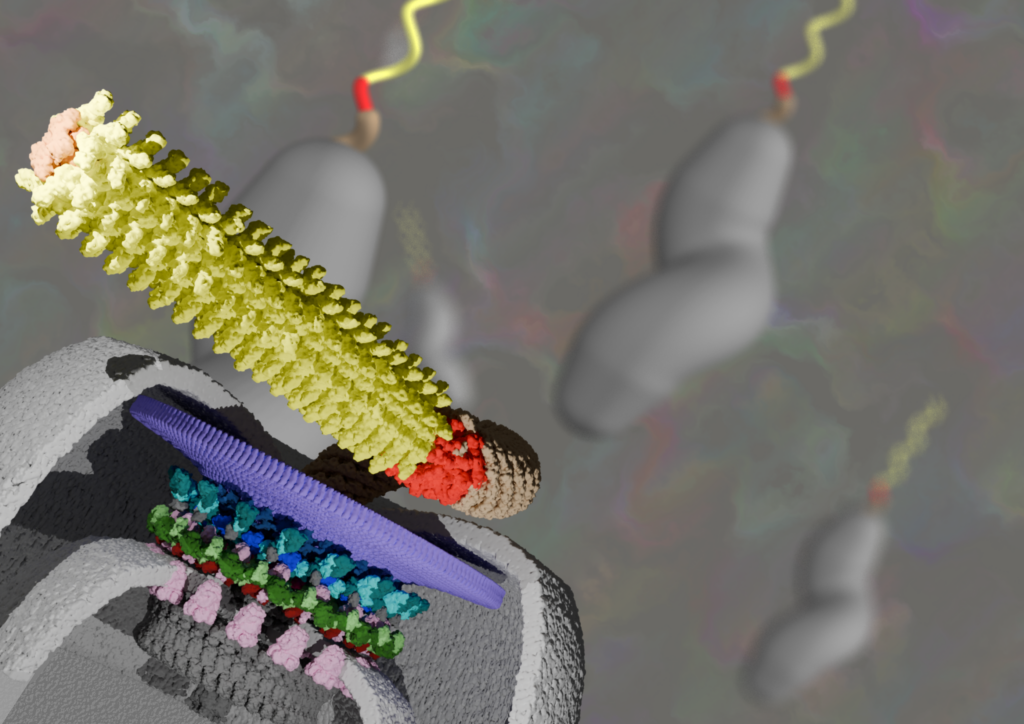
Electron cryo-microscopy of bacterial flagellar motors in situ reveals clues to their evolution
Bacterial flagellar motors are excellent case studies for understanding the mechanisms of molecular machines, and their evolution. Progress has been hampered by relatively low resolutions that fail to provide sufficient constraints for unambiguous molecular interpretation. Here I describe our recent work using electron cryo-microscopy to attain subnanometre resolutions of flagellar motors in situ, and the biological insights that this enables.

Nicholas Dale from the University of Warwick, UK
Following a degree in Zoology from Cambridge, Nicholas obtained a PhD in Neuroscience at the University of Bristol with Prof Alan Roberts. He was a post-doctoral fellow with Prof Sten Grillner at the Karolinska Institute, Stockholm, and then with Dr Eric Kandel at Columbia University, New York. He returned to the UK in 1989 to hold a Royal Society University Research Fellowship at Bristol. Nicholas then gained a Royal Society Locke Research Fellowship and held this initially at Bristol but moved to the University of St Andrews in 1995. In 2000, he moved to his current position as the Ted Pridgeon Professor of Neuroscience at the University of Warwick. In 2015, he gained a Royal Society Wolfson Research Merit Award for his work on CO2 chemosensing.
Talk Abstract
CO2–sensing via connexins: structural biology, physiology and evolution
Connexins canonically form gap junction channels that allow the passage of ions and small molecules between adjacent cells. However, they can also act as unopposed hemichannels, opening to the extracellular medium. Hemichannels of the three most widely expressed connexins in the human body are directly gated by CO2. Although CO2 is often regarded as a metabolic waste product, the ubiquity of CO2-sensitive connexin hemichannels suggests that CO2 may also be an important physiological signal. We have discovered that CO2 gates connexin hemichannels via a non-enzymatic carbamylation mechanism at an ancient evolutionarily conserved motif that is present in all CO2-sensitive connexins. I shall outline our understanding of how CO2 opens connexin channels and discuss some of the physiological/behavioural functions of connexin-mediated CO2 signalling. I will put forward our evidence for CO2acting as a specific cell-to-cell signal akin to a neurotransmitter. While the connexin gene is considered to be a chordate innovation, I will present our evidence that connexins have deep evolutionary origins from connexin-like domains in genes relating to innate immunity in Cnidaria and Protostomes.

Shannan Foylan from the University of Strathclyde, UK
Shannan is a postdoctoral research scientist at the Strathclyde Institute for Pharmacy and Biomedical Sciences, Glasgow UK. She has multidisciplinary research interests in molecular biology, transporter biology in disease and the optical design and implementation of state-of-the-art high and super-resolution optical imaging. Shannon completed her PhD in 2023 in the Department of Physics at the University of Strathclyde, developing several novel high throughput imaging methods with improved axial resolution for quicker imaging of larger cell populations. Her current research focuses on elucidating the mechanism for dispersal of the glucose transporter molecule GLUT4 in the plasma membrane of insulin responsive cells, with the long-term aim of understanding the impact of insulin-resistance/diabetes on this novel aspect of GLUT4 cell biology.
Talk Abstract
After a meal, rising blood glucose triggers insulin release from the pancreas, promoting glucose uptake into tissues via insulin-responsive molecular glucose transporters (GLUTs). In muscle and adipose cells, GLUT4 is the primary transporter responsible for this process, and defects in its regulation are hallmarks of insulin resistance and Type-2 diabetes. While intracellular trafficking of GLUT4 is well characterised, less is known about its organisation and diffusion within the plasma membrane (PM). Previous work has shown that GLUT4 forms stationary clusters under basal conditions but redistributes into diffusing monomers following insulin stimulation, a process requiring the scaffold protein EFR3.
To investigate GLUT4 nanoscale organisation, we applied both Single Molecule Localisation Microscopy and Ultrastructure Expansion Microscopy (U-ExM) to adipocytes and cardiomyocytes. We have established a pipeline for applying U-ExM to murine-derived adipocyte cell monolayers, achieving four-fold isotropic expansion and enabling large-volume, super-resolution imaging of GLUT4 using standard diffraction-limited systems. This approach provides an accessible alternative to costly super-resolution platforms for metabolic studies.
Complementary live-cell imaging and Fluorescence Correlation Spectroscopy revealed distinct dynamic behaviours of clustered versus dispersed GLUT4 molecules. Together, our findings uncover differences between basal and insulin-stimulated states and provide tools to study GLUT4 dynamics and insulin responsiveness in situ in metabolic tissues.

Stephan Gruber from the University of Lausanne, Switzerland
More information and talk abstract coming soon!

Sir Steve Jackson from the Cancer Research UK Cambridge Institute, UK
Sir Stephen (Steve) Jackson FRS, FMedsci, FAACR is a Professor of Biology at the University of Cambridge and Senior Group Leader at the CRUK Cambridge Institute (CI). After graduating in biochemistry from the University of Leeds, Steve carried out his PhD at Imperial College London and the University of Edinburgh, then carried out post-doctoral training at UC Berkeley. He returned to the UK in 1991 to set up his research group in what is now the Gurdon Institute, and moved to the CI in 2022. He has received various national and international prizes, and was awarded a knighthood for his services to innovation and research in 2023.
Talk Abstract
Cellular responses to DNA damage: mechanistic insights and medical implications
Steve’s academic research has identified key principles by which cells respond to and repair DNA damage and defined how their dysfunction yields cancer and other age-related diseases. In 1997, Steve founded the drug-discovery company KuDOS Pharmaceuticals, which developed and took into first patients the PARP inhibitor drug olaparib (LynparzaTM), now marketed worldwide for certain ovarian, breast, pancreatic and prostate cancers. In 2010, Steve founded Mission Therapeutics to exploit advances in protein ubiquitylation and deubiquitylation to derive new medicines. In 2018, he conceived of and co-founded Adrestia Therapeutics, which in 2023 was acquired by Insmed Inc.
His laboratory in the CI focuses on defining mechanisms by which cells detect, signal the presence of and repair DNA damage, and exploring how this knowledge might lead to new therapeutic approaches. This work involves biochemistry and precision cell biology underpinned by CRISPR-based genome engineering and genetic screens.

Alex Johnson from the University of Exeter, UK
More information and talk abstract coming soon!

Joergen Kornfeld from the MRC Laboratory of Molecular Biology, UK
More information and talk abstract coming soon!

Peijun Zhang from the University of Oxford, UK
Peijun Zhang is a Professor of Structural Biology in the Nuffield Department of Medicine at the University of Oxford and the founding director of eBIC (the UK National Electron Bio-imaging Centre) at the Diamond Light Source. She obtained her B.S. in Electrical Engineering and M.S. in Solid State Physics from Nanjing University, and Ph.D. in Biophysics and Physiology from University Virginia. She was a post-doctoral fellow and promoted to a staff scientist at the National Cancer Institute, NIH. She joined University of Pittsburgh School of Medicine as a principal investigator in 2006 and was subsequently granted tenure. In 2016, she moved to the University of Oxford and Diamond Light Source. She has received many prestigious grants, including a Wellcome Investigator Award, a Wellcome Discovery Award and an ERC Advanced Grant. She was elected a Member of the European Molecular Biology Organization.
Talk Abstract
Professor Peijun Zhang is a leading expert in cryo-electron microscopy (cryoEM) and cryo-electron tomography (cryoET), pioneering in situ structural biology approaches to visualize macromolecular complexes directly within their native cellular environments. Her research integrates method development with biological discovery to achieve a multi-scale, atomistic understanding of infection and cellular signalling mechanisms.
A central focus of her work is the structural biology of viral infection, particularly HIV-1 and other human viruses. Her group has developed and applied advanced correlative cryoEM/cryoET workflows—including cryoFIB/SEM for cellular lamella preparation and the high-resolution tomography software emClarity—to resolve macromolecular assemblies in native cells at near-atomic resolution. These innovations have enabled fundamental discoveries in HIV-1 nuclear entry, capsid–host interactions, viral maturation, and SARS-CoV-2 fusion and cytopathy, providing mechanistic insights that inform antiviral and vaccine strategies.
Beyond virology, Professor Zhang’s research extends to bacterial signalling systems, including chemotaxis pathways, using time-resolved cryoEM and cryoET to dissect dynamic macromolecular assemblies. Across all projects, her laboratory combines in situ structural analysis with high-resolution single particle cryoEM and complementary computational and biophysical/biochemical methods to bridge cellular context and molecular mechanism.


